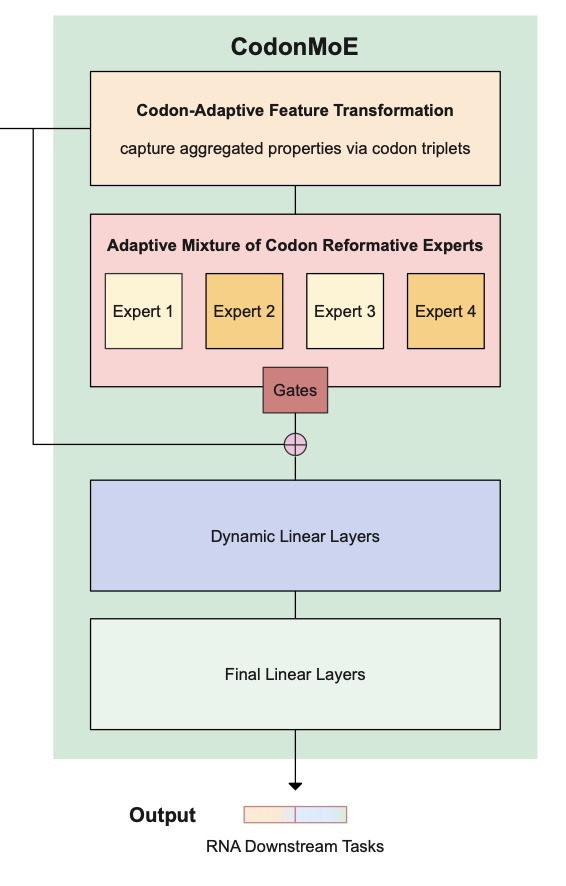
We introduce novel Adaptive Mixture of Codon Reformative Experts (CodonMoE) that can be incorporated into DNA gLMs in order to adapt them for mRNA-based predictive tasks. We show that, by using this plug-and-play operator, DNA-based gLMs can achieve performance similar to that of RNA-trained models on mRNA tasks. We further show that recent, efficient sub-quadratic DNA-based state space model (SSM) architectures can be used with the CodonMoE to achieve parameter- and computationally-efficient predictions for mRNA tasks. Specifically, experimental results demonstrate that CodonMoE improves diverse DNA-based backbones by a large margin, with some models achieving comparable or superior performance to current state-of-the-art RNA- specific models across several downstream tasks, while reducing both time complexity and model parameters. Our results provide a path for focusing development efforts of gLMs on DNA models, which can then be adapted to mRNA tasks. Because DNA data is more prevalent than assembled mRNA data, and modeling efforts can focus on a single class of model, this is likely to foster improved DNA models for mRNA tasks at lower computational cost and is a significant step towards unifying genomic language modeling.