Herbert A. Simon Professor of Computer Science
Ray and Stephanie Lane Computational Biology Department
Affiliate Faculty, Machine Learning Department
Director, Center for Machine Learning in Health
Director, Center for Innovation in Health
Co-Director, Joint CMU-Univ Pittsburgh Ph.D. Program in Computational Biology

Room 7719, Gates-Hillman ComplexContinue reading...
Carnegie Mellon University
5000 Forbes Ave
Pittsburgh, PA 15213

Current Group Members
- Dr. Guillaume Marçais, Senior Systems Scientist
- Shane Elder, CPCB Ph.D student
- Shiyi (Zoe) Du, CPCB Ph.D. student
- Jiayi Li, CPCB Ph.D. student
- Gün Kaynar, CPCB Ph.D. student
- Xander Song, CPCB Ph.D. student
- Yuxin Lu, CPCB Ph.D. student
- Hong-Sheng Lai, MSCB student
- Shijie Tang, MSCB student
- Zhaoyi You, MSCB student
Pretrained language models often rely on superficial features that appear predictive during training yet fail to generalize at test time, a phenomenon known as shortcut learning. Existing mitigation methods generally operate at training time and require heavy supervision such as access to the original training data or prior knowledge of shortcut type. We propose Shortcut Guardrail, a deployment-time framework that mitigates token-level shortcuts without access to the original training data or shortcut annotations. Our key insight is that gradient-based attribution on a biased model highlights shortcut tokens. Building on this finding, we train a lightweight LoRA-based debiasing module with a Masked Contrastive Learning (MaskCL) objective that encourages consistent representations with or without individual tokens. Across sentiment classification, toxicity detection, and natural language inference under both naturally occurring and controlled shortcuts, Shortcut Guardrail improves overall accuracy and worst-group accuracy over the unmitigated model under distribution shifts while preserving in-distribution performance.
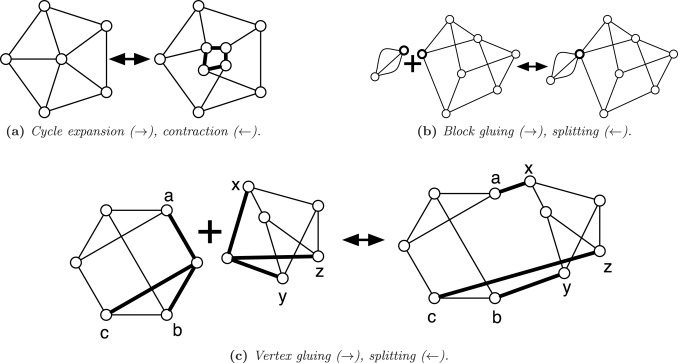
String algorithms make it possible to process, store, and manipulate text with computational efficiency, with applications ranging from search engines and social networks that regularly process terabytes of information to areas like genomics, where the genome of an organism can be encoded as a long string of letters. This book provides an incisive introduction to the concepts and applications that every practitioner in the field needs to know. Ideal for the classroom and self-study, it guides readers from the fundamentals of string processing to advanced computational methods, presenting useful data structures and proof techniques for strings and other data and serving as an on-ramp to doing cutting-edge research in string algorithms.

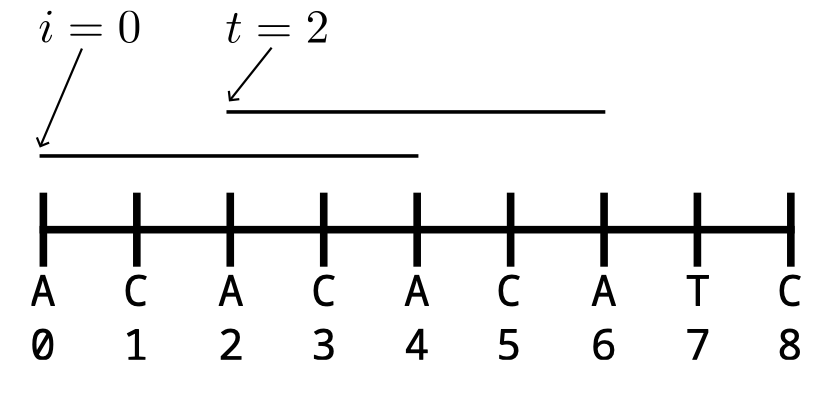
We study Turnpike with uncertain measurements: reconstructing a one-dimensional point set from an unlabeled multiset of pairwise distances under bounded noise and rounding. We give a combinatorial characterization of realizability via a multi-matching that labels interval indices by distinct distance values while satisfying all triangle equalities. This yields an ILP based on the triangle equality whose constraint structure depends only on the two-partition set {(r,s,t):yr+ys=yt} and a natural LP relaxation with {0,1}-coefficient constraints. Integral solutions certify realizability and output an explicit assignment matrix, enabling an assignment-first, regression-second pipeline for downstream coordinate estimation. Under bounded noise followed by rounding, we prove a deterministic separation condition under which y is recovered exactly, so the ILP/LP receives the same combinatorial input as in the noiseless case. Experiments illustrate integrality behavior and degradation outside the provable regime.
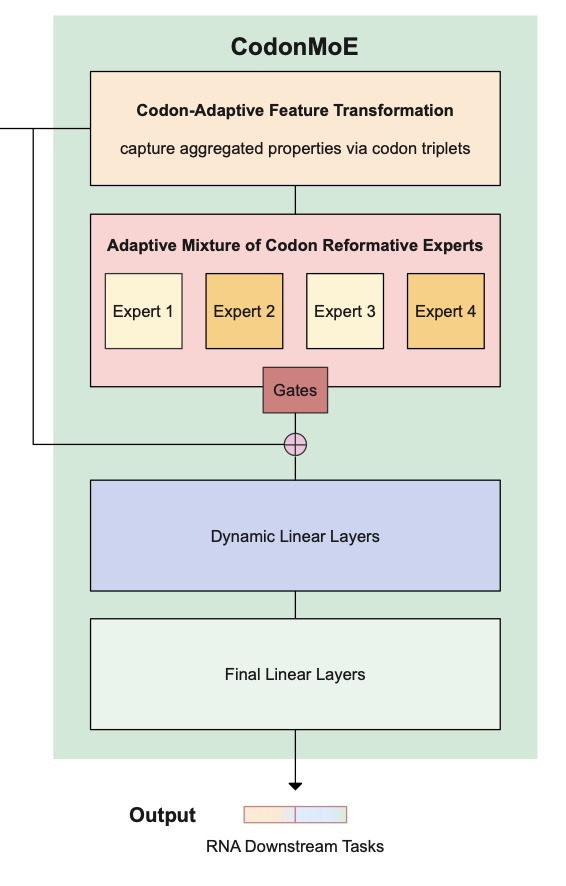
Optimizing synonymous codon sequences to improve translation efficiency, RNA stability, and compositional properties is challenging because the search space grows exponentially with protein length and objectives interact through long range RNA structure. Dynamic programming-based methods can provide strong solutions for fixed objective combinations but are difficult to extend to additional constraints. Deep generative models require large-scale, high-quality mRNA sequence datasets for training, limiting applicability when such data are scarce. Reinforcement learning naturally handles sequential decision-making but faces challenges in codon optimization due to delayed rewards, large action spaces, and expensive structural evaluation. We present CodonRL, a reinforcement learning framework that learns a structural prior for mRNA design from efficient folding feedback and demonstration-guided replay, and then enables user-controlled multi-objective trade-offs during inference.

Adverse events (AEs) resulting from medical interventions are significant contributors to patient morbidity, mortality, and healthcare costs. Prediction of these events using electronic health records (EHRs) can facilitate timely clinical interventions. However, effective prediction remains challenging due to severe class imbalance, missing labels, and the complexity of EHR records. Classical machine learning approaches frequently underperform due to insufficient representation of minority adverse event classes and limited capacity to capture interactions among patient demographics, administered medications, and associated complications. Methods: We introduce TASER-AE, a novel data augmentation pipeline tailored for structured EHR data, coupled with transformer-based classification. TASER-AE addresses these issues through an NLP-inspired data augmentation framework adapted for EHR, enabling effective minority-class representation in sparse and imbalanced clinical datasets.
Continue reading...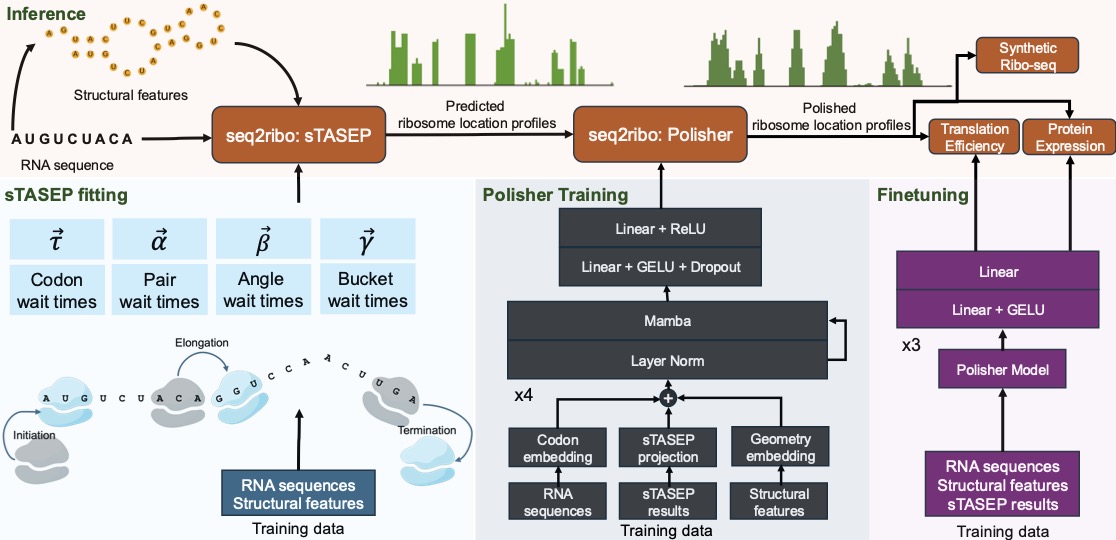
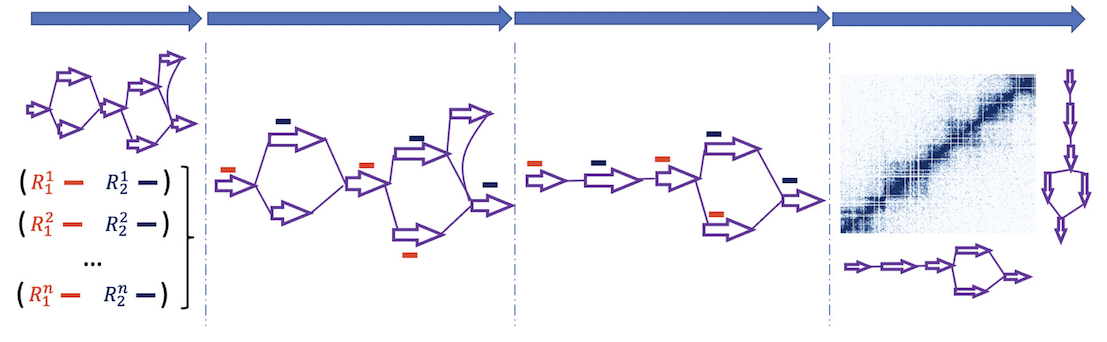
Motivation: Ribosome dynamics are vital in the process of protein expression. Current methods rely on ribosome profiling (Ribo-seq), RNA-seq profiles, and full genomic context. This restricts their use in de novo sequence design, like messenger RNA (mRNA) vaccines. Simulation-only approaches like the Totally Asymmetric Simple Exclusion Process (TASEP) oversimplify translation by focusing solely on codon elongation times.
Continue reading...Negative selection in the thymus limits autoimmunity by eliminating T cells that react strongly to self. Individual T cells, however, are only exposed to a small fraction of all self peptides during their “training” in the thymus, and it is puzzling how tolerance can be generalized to the remaining “test” self peptides across peripheral tissues in the body. Using a machine learning perspective, we show that such generalization is possible because the immune system satisfies two conditions: first that peptide abundance levels in the human thymus and periphery are highly correlated (i.e., training distribution ≈ test distribution), and second that cross-reactivity allows T cells to effectively learn binding information of similar peptides without explicitly interacting with all of them. Together, we show that sparse, random sampling of only 10% of self peptides in the thymus is sufficient to avoid reactivity to 90% of peripheral self, and we support this result with diverse experimental data. We then validate two predictions by our model; the first is that only 200–250 antigen presenting cells need to be seen by a T cell to ensure its robust selection, and the second relates how peptides missing from the thymus can drive auto-immunity of peripheral tissues. Overall, we provide a plausible answer to a long-standing question underlying adaptive immunity, and we highlight how generalization, a fundamental challenge faced by nearly every learning algorithm, is uniquely tackled by the immune system.